Dr. Fisk is an Associate professor in the Department of Chemistry at the University of Colorado Denver. He earned B.S. degrees with honors in Chemistry and Biochemistry from the Pennsylvania State University. Dr. Fisk completed his Ph.D. in Molecular Biophysics at the University of Wisconsin with Prof. Sam Gellman. At the University of Wisconsin, Dr. Fisk's research focused on understanding the fundamental forces involved in natural and non-natural polypeptide folding. Dr Fisk developed model systems for the study of parallel beta-sheet secondary structures in proteins, examined the folding of oligomers of non-natural beta amino acids primarily by NMR, and used phage display to identify new packing arrangemetns in protein coiled-coils. Dr. Fisk was an NIH postdoctoral research fellow at the California Institute of Technology in the laboratory of Prof. David. A. Tirrell. While at Caltech, Dr. Fisk worked on methodology to visualize newly synthesized proteins via the biosynthetic incorporation of reactive non-canonical amino acids.
Independent Career
“What I cannot create, I do not understand”
Richard Feynman’s profound statement guides all of the work in the Fisk laboratory. The ultimate test of understanding is our ability to design and synthesize systems that behave as predicted. The laboratory is motivated both by biomedical applications and a deep scientific interest in understanding the workings of natural systems. Synthetic biology requires the latter to do the former; engineering follows from a functionally complete quantitative understanding.
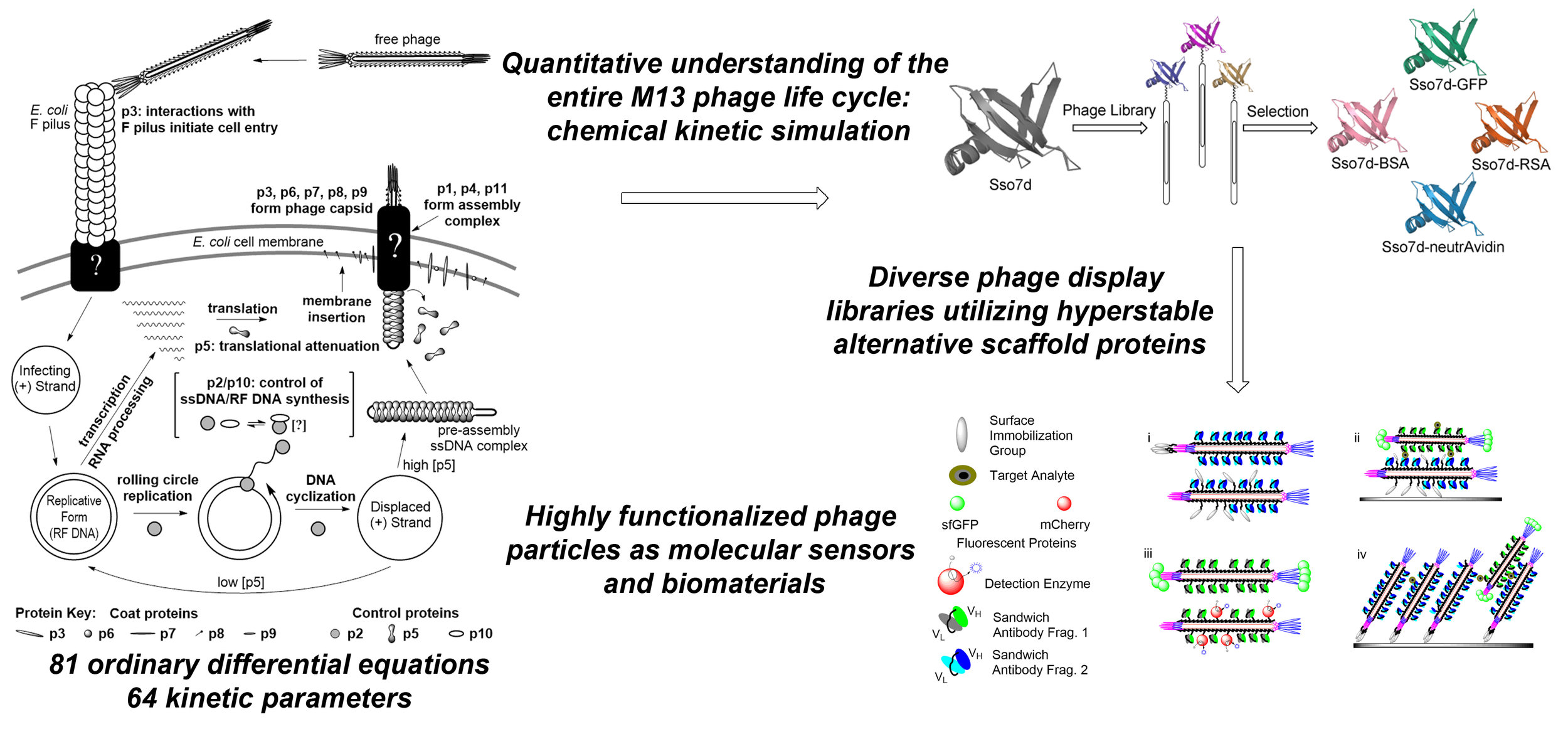
Three distinct research foci within the laboratory serve an overall desire to improve the understanding of complex biological systems to be able to fully exploit them for biotechnological purposes (e.g. diagnostics and therapeutics). Two research arms target engineered bacteriophage particles, mainly M13, as a molecular chassis for the development of new materials and devices. Phage systems are increasingly utilized in myriad biotechnological applications from batteries to biosensors. The nanoscale size and simple life cycles of bacteriophages make them ideal targets for rational redesign and engineering through synthetic biology. Our third research focus seeks to expand the genetic code by reassigning the meaning of sense codons. We are working to solve immediate problems in and challenges of both the phage display and non-canonical amino acid communities, but are working to do so in a way that knowledge gained and materials built for each research avenue leaves a legacy infrastructure that informs and enables future projects to produce more complex and functional systems. Ultimately, we hope to combine re-engineered bacteriophage systems with expanded genetic codes to build multivalent, multifunctional biomedical tools.